| |||||||||||||||||||||
Advanced AMDIS (Automated Mass Spectral Deconvolution and Identification System) from NIST has some important features. Coupling of mass spectra with retention indices: This is a topic which was unfortunately neglected in the past, but will be very important in the future for the identification of organic substances with GC-MS. Identification of substances only via mass spectra or retention indices (RI) may lead to false identifications. Mass spectra may be very similar and RI values may be false or overlapping. Its obvious that the more descriptors (vectors) exist for a certain compound, the more trustworthy the information will
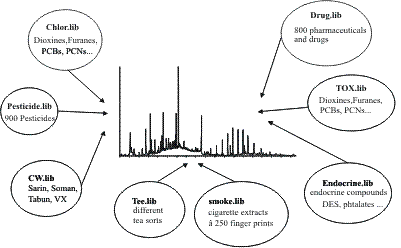
be. The concept of the AMDIS target libraries This ingenious topic is a milestone for scientists working with GC-MS or LC-MS. A stupid search for all substances existing in a certain chromatogram may be interesting but not truly clever. Always remember - CAS currently knows about 30 million substances (2002) - therefore a systematization is indispensable.
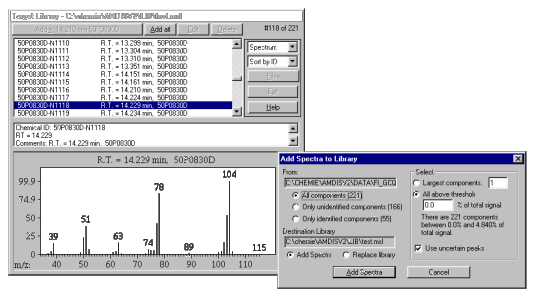
Library handling in AMDIS Many programs for GC-MS or LC-MS data processing can produce user (mass spectral) libraries from chromatograms. But AMDIS has a unique concept of handling these libraries. Peaks can be added not only peak by peak, but they can be processed in one step.
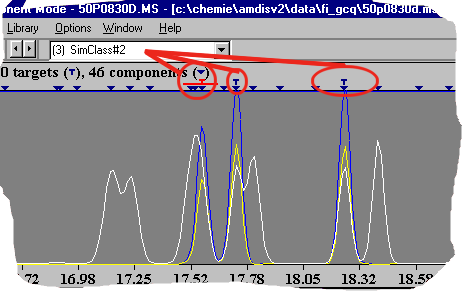
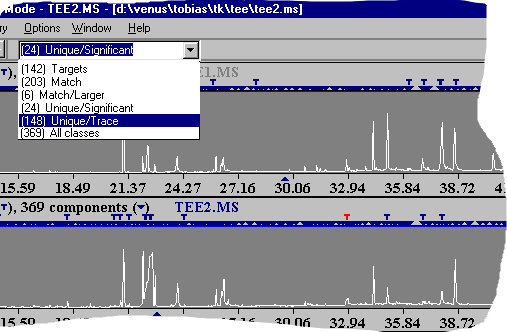
AMDIS post processing - spectral similarity This AMDIS topic classifies (if possible) all compounds in a data file. Even if database search fails you will get an information about certain substances. In most cases these compounds will be related structurally. In the picture you see a SIM-Class with all three dichlorobenzenes (marked with a T). In a former target library search they were not identified. But in the next step one can evaluate them en bloc.
This important step of systematization will fail in some cases (if the resolution is to low). But with the powerful library editor you can export all pure compounds and can classify the compounds with external programs. You will find some ideas in the amdis.net chemometrics section. AMDIS post processing - VARMUZA classifiers
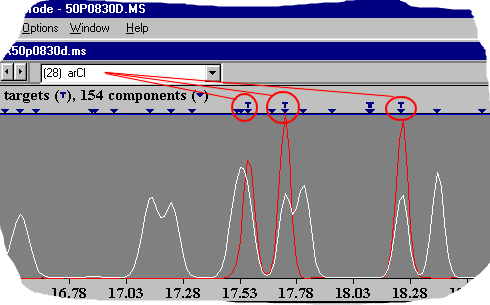
If you have no other information about a certain substance - you can choose this procedure. This is an expert algorithm which will detect special functional groups or substructures. You will find the substructures in the AMDIS handbook (amdis.net literature section).
As you can see in the graphics - all 3 dichlorobenzenes are correctly identified as an aromatic system with chlorine. AMDIS post processing - compare datafiles Every chromatographer knows this nice practice to overlay chromatograms and to compare them via the graphical peak pattern. This may be helpful in some cases but is only a very simple method.
|
[Home] [What is AMDIS?] [Chemometrics] [People] [External] [SiteMap] [About] |
© 2003 Tobias Kind - www.amdis.net - Alle Rechte vorbehalten. |